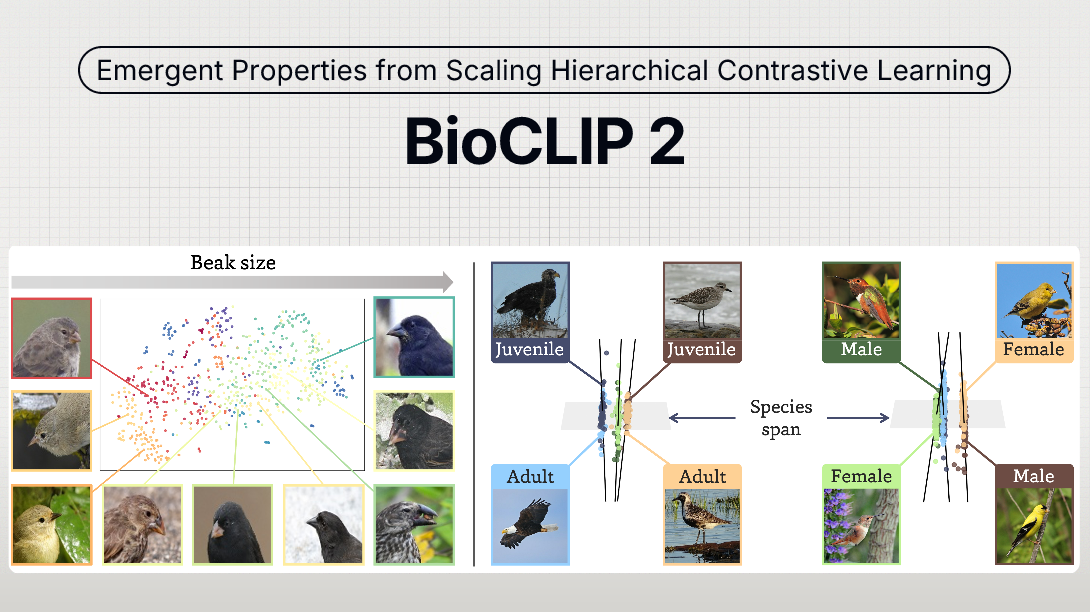
BioCLIP 2: Emergent Properties from Scaling Hierarchical Contrastive Learning
Foundation models trained at scale exhibit remarkable emergent behaviors, learning new capabilities beyond their initial training objectives. We find such emergent behaviors in biological vision models via large-scale contrastive vision-language training. To achieve this, we first curate TreeOfLife-200M, comprising 214 million images of living organisms, the largest and most diverse biological organism image dataset to date. We then train BioCLIP 2 on TreeOfLife-200M to distinguish different species. Despite the narrow training objective, BioCLIP 2 yields extraordinary accuracy when applied to various biological visual tasks such as habitat classification and trait prediction. We identify emergent properties in the learned embedding space of BioCLIP 2. At the inter-species level, the embedding distribution of different species aligns closely with functional and ecological meanings (e.g., beak sizes and habitats). At the intra-species level, instead of being diminished, the intra-species variations (e.g., life stages and sexes) are preserved and better separated in subspaces orthogonal to inter-species distinctions. We provide formal proof and analyses to explain why hierarchical supervision and contrastive objectives encourage these emergent properties. Crucially, our results reveal that these properties become increasingly significant with larger-scale training data, leading to a biologically meaningful embedding space.