BioCLIP 2.5 Huge
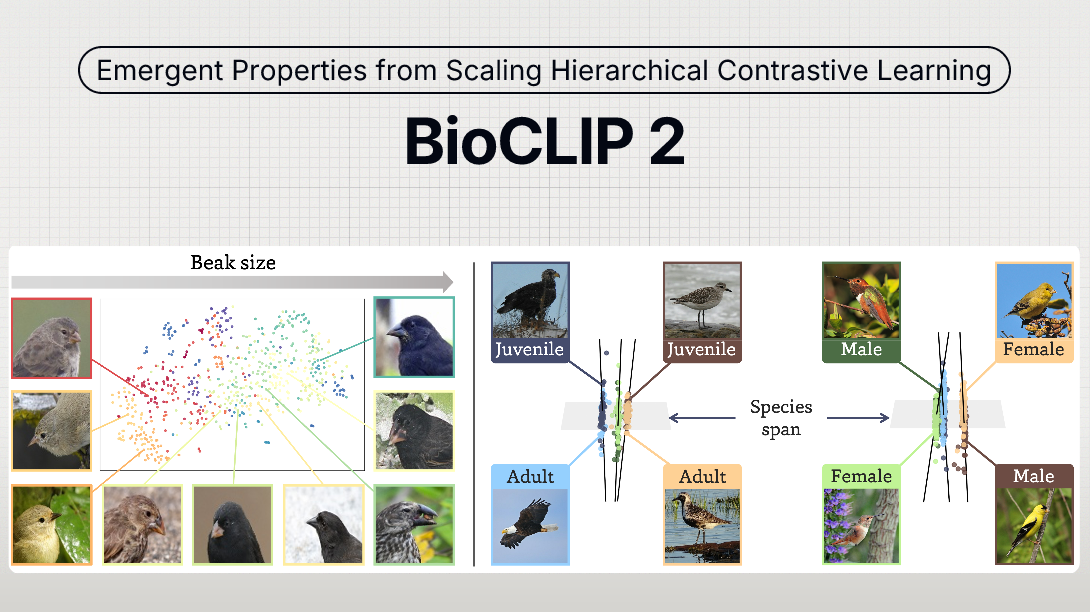
The largest model in the BioCLIP collection, BioCLIP 2.5 Huge was trained on 233-million images across more than 950-thousand taxa. Training was accelerated through an updated version of the BioCLIP 2 repository (v2.0.0). BioCLIP 2.5 exhibits emergent properties, extending beyond simple classification, to distinguish between life stages, sexes, and align embeddings with ecological traits like beak size.
The model achieves new state-of-the-art performance on both species classification and broader biological visual tasks, surpassing BioCLIP 2 by 5.7% and 3.5%, respectively. Especially in FishNet, which requires the model to distinguish different habitats, BioCLIP 2.5 Huge demonstrates a 8.7% performance improvement over BioCLIP 2.
- Architecture: ViT-H/14 (Huge)
- Training Data: 233 Million images (TreeOfLife-200M)
- Best for: High-accuracy tasks, fine-grained trait analysis, zero-shot learning, no inference speed or memory constraints.